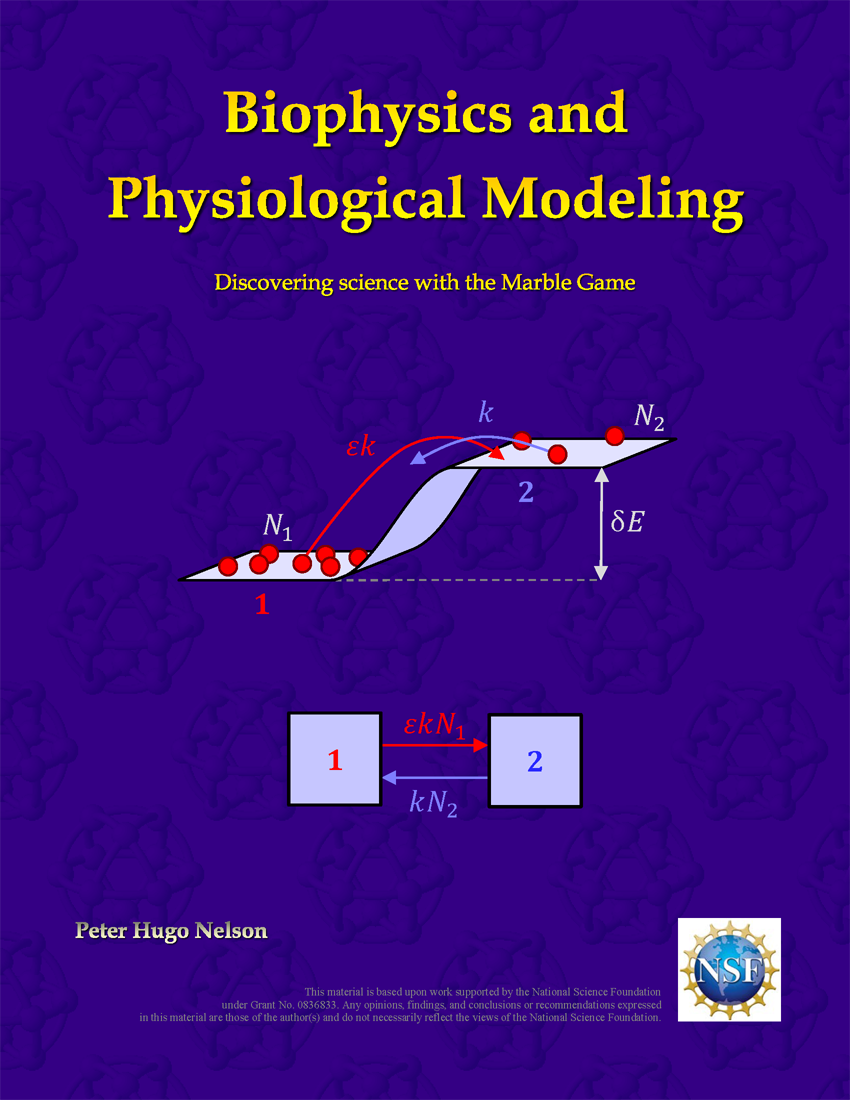
 Biophysics and Physiological Modeling
Biophysics and Physiological Modeling
| Free sample chapters | Videos | Instructor guides | Home |
There is a growing movement within biology to transform the undergraduate curriculum. Major goals are to develop active-learning teaching materials based on educational research, and to engage students in discovery-based activities so that they perceive biology as an evidence-based science with testable hypotheses that are supported by experimental data. As part of a National Science Foundation (NSF) grant, I'm developing a series of modules (now sample chapters for a book due to be published by Cambridge University Press) that engage students in inquiry-based activities to introduce them to the process of scientific modeling and validation. The chapters are a self-contained series of interactive self-study guides for undergraduates (both with and without calculus). The chapters are designed so that students can complete them without additional assistance, making them suitable for use as stand-alone homework exercises or computer labs, or for a complete discovery-based active-learning course.
These materials are aimed at biology, pre-med and physical science majors interested in learning about physiological systems and processes.
 Chapter 1 introduces the marble game and provides a gentle introduction to quantitative modeling using Excel. This marble game provides a realistic conceptual framework for understanding a wide range of molecular transport processes in physiology. These transport phenomena are so fundamental to physiology that "Transport Processes" was ranked as the second most important topic in pre-medical education (after "Nucleic Acids") in a 2011 survey conducted by the Association of American Medical Colleges (AAMC). In a 2015 IPLS report, the American Association of Physics Teachers (AAPT), identified transport processes as a crucial topic that is frequently neglected in introductory physics. Chapters 1, 2 and 3 have been peer reviewed by the Archive Board of the American Physiological Society (APS).
Chapter 1 was selected to be one of four resources featured on the front page of their NSF-funded online portal: "Life Science Teaching Resource Community" in November 2011 and Chapter 2 was a featured "Inquiry-Based K-12 Science" resource in March and April 2012.
If you’re interested in trying out Chapter 1, check out the LifeSciTRC listing, the Chapters homepage or go straight to Chapter 1.
Chapter 1 introduces the marble game and provides a gentle introduction to quantitative modeling using Excel. This marble game provides a realistic conceptual framework for understanding a wide range of molecular transport processes in physiology. These transport phenomena are so fundamental to physiology that "Transport Processes" was ranked as the second most important topic in pre-medical education (after "Nucleic Acids") in a 2011 survey conducted by the Association of American Medical Colleges (AAMC). In a 2015 IPLS report, the American Association of Physics Teachers (AAPT), identified transport processes as a crucial topic that is frequently neglected in introductory physics. Chapters 1, 2 and 3 have been peer reviewed by the Archive Board of the American Physiological Society (APS).
Chapter 1 was selected to be one of four resources featured on the front page of their NSF-funded online portal: "Life Science Teaching Resource Community" in November 2011 and Chapter 2 was a featured "Inquiry-Based K-12 Science" resource in March and April 2012.
If you’re interested in trying out Chapter 1, check out the LifeSciTRC listing, the Chapters homepage or go straight to Chapter 1.
In later chapters, we’ll use the marble game to develop simple models that can be implemented in an Excel spreadsheet to discover more about these transport process. Using the spreadsheet, we’ll discover what the practical implications of the model are, and then we’ll compare the model predictions with experimental data (if available). For example, in the early chapters, we’ll develop a simple (pharmacokinetic) model of how drugs such as acetaminophen (paracetamol) and penicillin are eliminated from the body. While this model (and the other models we’ll develop) are particularly simple (and are therefore easy to understand), the molecular modeling approach is actually a cutting-edge research technique! - a kinetic Monte Carlo simulation.
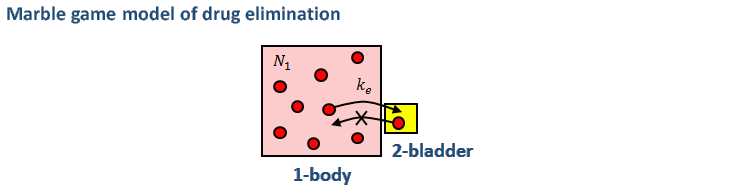
Once we’ve developed the drug elimination model, we’ll compare it's predictions with published data for elimination of piperacillin (a form of penecillin) by healthy volunteers and intensive care patients on heart-lung machines. As we'll discover the marble game model does a fantastic job of predicting what will happen in the healthy volunteers... There are numerous other topics that we can investigate with a similar approach including:
|
|
I’m currently working on developing new topics and approaches. If there is a topic that you’re interested in, that you think might be fun to do, please let me know.
-
Beta testing and evaluation
-
Chapters 1-3 have been peer-reviewed by the Archive Board of the American Physiological Society (APS). If you’re interested in trying out chapter 1, check out the LifeSciTRC listing, the Sample Chapters homepage or go straight to Chapter 1.
Addtional chapters are also available. If you’re interested in evaluating them, please email me. I'm also looking for collaborators who are interested in evaluating these materials as an example of how science education projects evolve and how to determine student learning benefits from curricular changes. If you are interested in collaborating on a Research Coordination Networks - Targeted Undergraduate Biology Education track (RCN-UBE) proposal, please let me know.
-
Self-study guides
- The chapters are self-contained workbooks. Hence, you (or your teaching assistant (TA)) can use them as supplemental homework assignments or computer labs, or for an entire discovery-based active-learning course.
-
Prerequisites and level
- The sample chapters assume high school chemistry, and (algebra-based) physics. No calculus required! However, optional sections are included for those students with calculus. They are also suitable as an introduction to scientific modeling and validation for students who didn’t take a formal course in biophysics, e.g. graduate or medical students in interdisciplinary Biophysics or Molecular Biology PhD programs or MD/PhD programs.
The physiological models developed in the sample chapters provide descriptive explanations that are very accessible to high school students (even if the chapters are not utilized directly). Hence, after working through the chapters, high school instructors can use the marble game conceptual framework to correctly describe biophysical phenomena in using principles that are accessible to high school students. This can be contrasted with traditional biophysical arguments that rely on thermodynamic principles (such as entropy, free energy, and chemical potential, which are not well-understood by high school students or even most undergraduates!). The marble game approach relies on simple kinetic reasoning that can be used by students to construct thermodynamically-sound mental models of processes such as diffusion and osmosis that can be presented to high school students without requiring the mathematical detail presented in the sample chapters.
Notwithstanding that the materials are accessible to well-prepared high school students; they also contain materials that have traditionally been taught at the graduate level. As an example, the kMC simulation technique (or the chemical master equation) is not in the traditional undergraduate curriculum even for chemistry or physics majors. The primary focus of the chapters is quantitative scientific modeling with physiology applications.
-
Background
- BIO 2010 - Transforming Undergraduate Education for Future Research Biologists (National Academies Press 2003) motivated me to write the NSF CCLI (now TUES) proposal for this project. The chapters also address many of the core competencies identified in the report of the AAMC-HHMI Committee: Scientific Foundations for Future Physicians published by the Association of American Medical Colleges (AAMC) 2009 and they address many of the issues discussed by the "Overarching and unifying key concepts and competencies - Group B - Quantitative Competency" working group at the AAAS conference: Transforming Undergraduate Biology Education: Mobilizing the Community for Change (July 15-19, 2009) sponsored by the NSF. My presentation at this conference was about this Biophysics and Physiological Modeling project. The revised Vision and Change Final Report is now available as are the Ratings of the Importance of Natural Sciences, Research Methods, and Statistics Topics on the MR5 Content Surveys, 5th Comprehensive Review of the Medical College Admission Test® (MCAT), MR5 Preliminary Recommendations, Creating Scientifically Literate Physicians, MR5 Science Content Survey of Undergraduate Institutions, Preview Guide for MCAT 2015 and MCAT website.
-
Student researchers/testers
- I would like to thank my students in
 Phys 111/112 - "Introductory Physics for the Life Sciences", Biol 310 - "Physiological Modeling", and Biol/Chem/Phys 323 - "Biophysics",
for their patience, support, encouragement and enthusiasm. I couldn’t have done it without you!
Phys 111/112 - "Introductory Physics for the Life Sciences", Biol 310 - "Physiological Modeling", and Biol/Chem/Phys 323 - "Biophysics",
for their patience, support, encouragement and enthusiasm. I couldn’t have done it without you!
I would also like to specifically thank the following students for their assistance: Nathan Kucera, Jonathan Rink, Ashley Rink, Stephanie Denny, David Faber, Matthew Harrison, James Norris, Muzamil Arshad, Amar Khatri, Luke Cox, Rama Wahood, Tricia Avanzado, Joy Holowicki, Madhish Patel, Jacob Polaski, Sabrina Sanchez, Alexis Wadowski, Mohammad Arfeen, Kristine Baldwin, Joseph Esner, Branko Milicevic, Daniela Janevska, David
Kamber, Robert Kasper, André Molina, Garrick Moll, Ankit Parikh, Jonathan Pollock, John Romal,
Mohammad Arfeen, Kristine Baldwin, Joseph Esner, Branko Milicevic, Daniela Janevska, David
Kamber, Robert Kasper, André Molina, Garrick Moll, Ankit Parikh, Jonathan Pollock, John Romal,  Sawyer Masonjones, Edyta Frankiewicz, Katie Maezes, Justin Villanueva, Tiffany Warzecha, Nicole Worthem, Thymur Chaudry, Owais Hameed, Robert Horsley, Joey Hua, Tahreem Iqbal, Bohdan Khomtchouk, Raquel Tobar Rubin, Megan Wisniewski, Maria Radcliffe, Dima Saigh, Amanda Stachowicz, Lina Savickas, Jared St. John, Diyanka Patel, Le-Ander Gibbs, Dana Cairns, Amy Klingbiel, Spencer Havis, Ian Martinek, Rex Rajan, Matt Raub, Bridgett Wrobel, Mario Mendez, Jeremy Bingen, Zahra Rehman,
Sawyer Masonjones, Edyta Frankiewicz, Katie Maezes, Justin Villanueva, Tiffany Warzecha, Nicole Worthem, Thymur Chaudry, Owais Hameed, Robert Horsley, Joey Hua, Tahreem Iqbal, Bohdan Khomtchouk, Raquel Tobar Rubin, Megan Wisniewski, Maria Radcliffe, Dima Saigh, Amanda Stachowicz, Lina Savickas, Jared St. John, Diyanka Patel, Le-Ander Gibbs, Dana Cairns, Amy Klingbiel, Spencer Havis, Ian Martinek, Rex Rajan, Matt Raub, Bridgett Wrobel, Mario Mendez, Jeremy Bingen, Zahra Rehman, Julie Carroll, Humdoon Choudhry, Rabia Choudhry, Nick Gill, Seerat Hassan, Maggie Julien, Nemanja Krstic,
 Kiran Munir, Zach Oesterreicher, Cameron Pombert, Mirela Trifonova, Martin Hirsch, Maryam Moeed, Jozef Palasiewicz, Kelsey Zimmermann, Jonathan Bell, Nathan Kuhlmann, Chand Bhanot, Genice Onwuka, Anna Lichtiger, Laura Adair, Steven Kerr, Abbie Horchar, Korbin LeClair, Nikka Hubert, Corey Proctor, Lucretia Deam, Emily Bradford, Michaela Allred, Sarah Sedaghat, Sam Schopler, Makayla Carver, Charles Bookheimer, Caleb Amstutz, Kaylee Zabava, Nina Palamaris, Evan Rhodes, Maitha Ali, Fatemeh Agha Mohammadi, Ryan Hill, Colby Hinkle, Andrea Muniz-Alvarado, Jasmine Greenwood, Alli Jackson, Sara Clodfelter, Danielle Galipeau, Elizabeth Ruprecht, Kylie Richardson, Elijah Paul Gregory, Exel Valle Estrada, Chavis Junior, Kelsey Wilson, Jasmine Voss, Jasmin Cuspard,
Thaddeus Smith, Marcel Edinbugh, Oreoluwa Owoseeni, Collins Ijale, Onyinyechi Agbo, Zipporah Davis, Dy'Mond Smith, Nevaeh Goins, Taiyonni Hill, Elizabeth Olajide, Ngozi Nwankwo, Mya Robinson, Jordan Lowther, Malia Jennings, Madyson Walker, Odinakachukwu Agubuzo, Grace Aruleba, Ada'Jiah Burton, Angele Hill, Emma Williams, Gabrielle White, Ryan Biddle, Kendall Wilson, Arianna Harris, and Jahneyah Boyce.
Kiran Munir, Zach Oesterreicher, Cameron Pombert, Mirela Trifonova, Martin Hirsch, Maryam Moeed, Jozef Palasiewicz, Kelsey Zimmermann, Jonathan Bell, Nathan Kuhlmann, Chand Bhanot, Genice Onwuka, Anna Lichtiger, Laura Adair, Steven Kerr, Abbie Horchar, Korbin LeClair, Nikka Hubert, Corey Proctor, Lucretia Deam, Emily Bradford, Michaela Allred, Sarah Sedaghat, Sam Schopler, Makayla Carver, Charles Bookheimer, Caleb Amstutz, Kaylee Zabava, Nina Palamaris, Evan Rhodes, Maitha Ali, Fatemeh Agha Mohammadi, Ryan Hill, Colby Hinkle, Andrea Muniz-Alvarado, Jasmine Greenwood, Alli Jackson, Sara Clodfelter, Danielle Galipeau, Elizabeth Ruprecht, Kylie Richardson, Elijah Paul Gregory, Exel Valle Estrada, Chavis Junior, Kelsey Wilson, Jasmine Voss, Jasmin Cuspard,
Thaddeus Smith, Marcel Edinbugh, Oreoluwa Owoseeni, Collins Ijale, Onyinyechi Agbo, Zipporah Davis, Dy'Mond Smith, Nevaeh Goins, Taiyonni Hill, Elizabeth Olajide, Ngozi Nwankwo, Mya Robinson, Jordan Lowther, Malia Jennings, Madyson Walker, Odinakachukwu Agubuzo, Grace Aruleba, Ada'Jiah Burton, Angele Hill, Emma Williams, Gabrielle White, Ryan Biddle, Kendall Wilson, Arianna Harris, and Jahneyah Boyce.

- P. H. Nelson Biophysics and Physiological Modeling: Model Validation and Penicillin. (2010) MedEdPORTAL: https://www.mededportal.org/publication/8081, https://doi.org/10.15766/mep_2374-8265.8081 (part of the uScience Inaugural Collection).
- P. H. Nelson A permeation theory for single-file ion channels: One- and two-step models (2011) J. Chem. Phys. 134, 165102*
(a chapter on ion channel permeation is in preparation) - P. H. Nelson Teaching Physiology with the Marble Game (2011) lifescitrc.org http://www.lifescitrc.org/resource.cfm?submissionID=4999
(Featured resource in November 2011)
(Featured Inquiry-Based K-12 Science resource in March and April 2012) - P. H. Nelson Teaching Physiology with the Marble Game 2: Algorithms and TYLENOL® (2012) lifescitrc.org http://www.lifescitrc.org/resource.cfm?submissionID=6152
- P. H. Nelson Teaching Physiology with the Marble Game 3: Finite difference method & oxygen (2013) lifescitrc.org http://www.lifescitrc.org/resource.cfm?submissionID=7623
- P. H. Nelson Osmosis and thermodynamics explained by solute blocking (2017) Eur. Biophys. J. 46, 59-64
- P. H. Nelson Introductory models of COVID-19 in the United States (2021) The Biophysicist 2 (3), 74-98.

- P. H. Nelson (2011). Connecting ion channel simulations to experiment APS March Meeting 2011 56(1)
- P. H. Nelson (2011). Teaching Physiology with the Marble Game FASEB J. 25, 481.5
- S. I. Sanchez, A. Wadowski and P. H. Nelson (2011). Developing and testing a quantitative model of ion permeation based on the MacKinnon mechanism FASEB J. 25, 1042.24
- A. Wadowski, S. I. Sanchez and P. H. Nelson (2011). A mathematical equivalence between the MacKinnon model and the association/dissociation model explains its apparent success in explaining ion channel permeation FASEB J. 25, 1042.25
- P. H. Nelson (2012). Teaching Biophysics with the Marble Game Biophysical Society 56th Annual Meeting 2012 Pos-L333
- P. H. Nelson (2012). A new pedagogy for ion channels: they’re permeases, not pores! FASEB J. 26, 719.9
- P. H. Nelson (2013). Teaching Outside the Box and Understanding Biophysics Using Microsoft Excel - 2013 TUES Principal Investigators (PIs) Conference A4 and 30
- P. H. Nelson (2013). Teaching Introductory STEM with the Marble Game Biophysical Society 57th Annual Meeting 2013 2732-Pos
- P. H. Nelson (2013). A new pathway to STEM - the marble game FASEB J. 27, lb887
- P. H. Nelson (2014). Biophysics in the Undergraduate Curriculum Biophysical Society 58th Annual Meeting 2014 1099-Pos
- P. H. Nelson (2014). Conference on Introductory Physics for the Life Sciences, Arlington, VA, 2014 Biophysics in the Undergraduate Curriculum
- P. H. Nelson (2015). Osmosis, colligative properties, entropy, free energy and the chemical potential Updates and Resources for Introductory Physics for Life Science II AAPT Winter Meeting, San Diego, CA, 2015 GH03
- P. H. Nelson (2015). Single-File Water Permeation through Aquaporin Channels Biophysical Society 59th Annual Meeting 2015 2204-Pos
- P. H. Nelson (2015). Teaching Science Like We Do Science: Integrating Research and Education Workshop Invited presentation titled "Keep Reading and Do Science: A diffusive model of osmosis" Biophysical Society 59th Annual Meeting 2015
- P. H. Nelson (2021). Modeling the Coronavirus Pandemic in The United States Teaching the Introductory Physics for the Life Sciences (IPLS) Course AAPT Virtual Winter Meeting, 2021 C1.1-02
- P. H. Nelson (2021). IPLS – The Physics That Life-Science Students Want and Need Teaching the Introductory Physics for the Life Sciences (IPLS) Course AAPT Virtual Winter Meeting, 2021 C1.1-04 YouTube Video
- P. H. Nelson (2021). Introductory Models of the Covid-19 Pandemic in the United States Biophysical Society 65th Annual Meeting 2021 YouTube Video
- P. H. Nelson (2021). Incorporating Molecular Biophysics in the Undergraduate Curriculum Moving beyond listening in BMB education Spotlight Session ASBMB Experimental Biology 2021 April 27-30 R2784 YouTube Video
- P. H. Nelson (2022). Biophysics in the Undergraduate Curriculum Biophysics in the 21st Century Curriculum I, AAPT Virtual Winter Meeting, 2022 A3.1-03
- P. H. Nelson (2022). Playing the marble game in Excel Introductory Courses I AAPT Summer Meeting, Grand Rapids, Michigan, July 9 - 13, 2022, HB06
- Thaddeus Smith and P. H. Nelson (2022). Modeling the Omicron Wave of COVID-19 with the SIR Model - Award for Best Poster SPS Undergraduate Posters AAPT Summer Meeting, Grand Rapids, Michigan, July 9 - 13, 2022, SPS01
- P. H. Nelson (2024). Biophysics and computational modeling in the undergraduate curriculum Biophysical Society 68th Annual Meeting February 10–14, 2024, Philadelphia, Pennsylvania, USA
- P. H. Nelson (2024). Computation in the IPLS Course Examining the Introductory Physics for the Life Sciences (IPLS) Course AAPT Summer Meeting, Boston, Massachusetts, July 6 - 10, 2024, GC-03
- P. H. Nelson (2024). Computation in the IPLS Course IPLS Poster Roundtable AAPT Summer Meeting, Boston, Massachusetts, July 6 - 10, 2024, FC-01
-
Funding and awards
-
Six Star Science Teacher-Recommended Collection: Cellular Transport
NSF Grant DUE-0836833
Transforming Undergraduate Education in STEM: Building a Community to Transform Undergraduate STEM Education - 2013 TUES Principal Investigators (PIs) Conference
AAAS Conference: Transforming Undergraduate Biology Abstracts
AAMC MedEdPORTAL for Undergraduate Science (uScience)
HHMI Grant 52002669
Title III Grant - Benedictine University
NIH Post-Doctoral Fellowship in Quantitative Biology GM20584
-
Searching, resources and portals
-
Physics of Life (2022) (National Academies of Sciences, Engineering, and Medicine) - explains why this book is needed,
NSDL, ben (BiosciEdNet), www.lifescitrc.org, CSERD, the Gateway, comPADRE, IPLS Final Report, www.the-aps.org, Disseminating CCLI Educational Innovations, SALG, URSSA, Mobilizing STEM Education for a Sustainable Future, ESTEEM, PER User's Guide, NEXUS, Websites, DiscoverEd, Open Courseware, HHMI's BioInteractive, Biophysical Mechanisms (Biophysics Textbook Online - Biophysical Society), HHMI Membrane Awakening, BPM Hotlist
 Circle4.com
Circle4.comThis material is based upon work supported by the National Science Foundation under Grant Nos. 0836833, 1817282, and 2306506.
Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
This page was last updated on March 4, 2025 by pHn.
© Peter Hugo Nelson 2009-2025
All Rights Reserved
Nelson, P. H. (2023) Biophysics and Physiological Modeling; Available from: http://circle4.com/biophysics/
Cite an individual chapter (e.g. Chapter 1) as:
Nelson, P. H. (2023) Biophysics and Physiological Modeling - Chapter 1: Introduction - marble game (v.4.6); Available from: http://circle4.com/biophysics/
- Copyright Information:
- * This article appeared in J. Chem. Phys. and may be found at (P. H. Nelson J. Chem. Phys. 134, 165102 (2011)). Copyright American Institute of Physics. This article may be downloaded for personal use only. Any other use requires prior permission of the author and the American Institute of Physics.
